Speaking a thousand words – how a cancer image collection is set to improve AI diagnosis
Artificial intelligence is set to revolutionise many aspects of cancer diagnosis and treatment – but an AI is only as good as the data used to train...
Commercial data partnerships are essential to maximising the impact of Patient-Derived Data but must be established in a safe, secure and transparent manner. To ensure this we have developed, in close consultation with patients, a set of guiding principles for commercial data partnerships that protect patients, our partners and our data.
Over the past five to 10 years, thanks to advancements in data generation and analytical technologies, the role of data in tackling cancer has become increasingly important.
Cancer Research UK is the leading non-governmental funder of cancer research in the world. This means our research network is responsible for the generation of huge amounts of data generated through clinical trials or research projects performed outside standard clinical care (Patient-Derived Data). At Cancer Research Horizons, we believe that through establishing commercial data partnerships, we can maximise the impact of this Patient-Derived Data and drive further patient benefit.
We recognise the sensitivities involved in sharing of Patient-Derived Data, and are committed to maintaining trust, transparency and accountability in our data partnerships. That's why, in consultation with patients, we've developed our 'Guiding Principles for Commercial Data Partnerships'.
‘Data partnerships’ are collaborations, licences and other forms of formal partnerships with a commercial entity that relate specifically to Patient-Derived Data in which the commercial entity acquires rights or access to the underlying Patient-Derived Data.
We conducted several consultations with people affected by cancer during development of these principles. From these consultations we wanted to understand
1) how to effectively involve patients in decisions around commercial data partnerships
2) the principles we should adopt to ensure any partnerships involving Patient-Derived Data have clear benefits for people affected by cancer without unacceptable risk.
Patient involvement formed the bedrock from which our guiding principles were built in addition to public and private sector consultation, ensuring they represent emerging best practice. These principles will be reviewed and updated in consultation with patients at least annually to ensure that they are up to date and reflect the evolving landscape.
Our guiding principles form just one part of our overarching framework for ensuring commercial data partnerships are in the interests of people affected by cancer. In addition to these principles, Cancer Research Horizons carries out detailed due diligence, involve people affected by cancer and enforce legal contracts before any commercial organisation accesses patient-derived data.
We are committed to establishing transparent commercial data partnerships. We are working toward generating summaries of each partnership we have entered since the launch of our Commercial Data Partnership Guiding Principles in July 2022. These can be found on the list below, which will be updated constantly. The outputs of these collaborations are discussed with our patient involvement group and used to inform the evolution of our guiding principles.
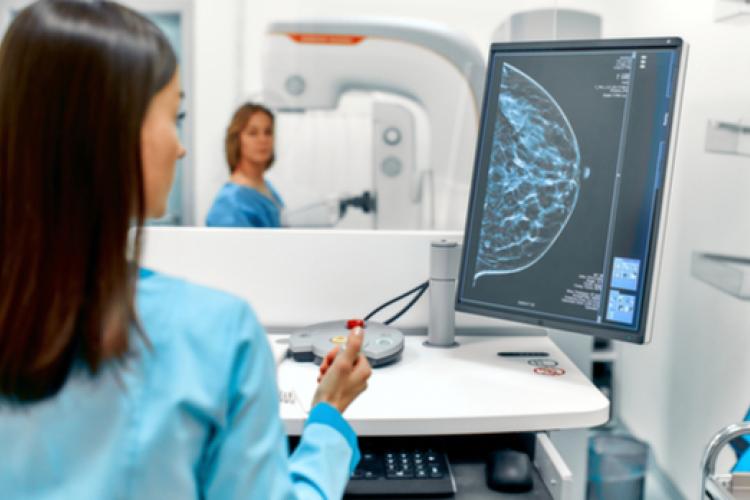
Mammograms (breast X-rays) are a crucial part of breast cancer screening, implemented across the world as part of national breast screening initiatives. These scans are normally analysed by radiologists, but the quantity and complexity of these images means that additional tools are needed to support their interpretation. Google Health is developing tools to support the detection of breast cancer in mammograms and this collaboration provides Google with access to de-identified data from the OPTIMAM study to help support the development and validation of these tools.
ScreenPoint Medical develops artificial intelligence software that helps radiologists to detect breast cancer earlier and faster. This software works as an assistant to the radiologist by analysing the images to look for signs or risk of breast cancer. To ensure equal and fair support to all women, in particular those attending the UK breast cancer screening program, de-identified data from patients participating in the OPTIMAM study are used in the development and validation of these solutions.
Genialis has developed a proprietary machine learning framework which models disease biology with the goal to inform treatment decisions and realise personalised medicines for patients. In supporting the development of these biomarker algorithms, the company relies on access to diverse datasets to ensure the biomarkers are accurate and robust across different demographics. This collaboration provides Genialis access to a subset of S:CORT data to support the development and validation of the next generation of RNA based precision medicine solutions
Cancer is a heterogenous disease with distinct characteristics in every patient. These differences are largely due to the random nature of genetic mutations which allow uncontrolled growth in cancer cells, thus highlighting the importance of effectively selecting patient populations on the basis of their predicted sensitivity to specific drugs. CDC7 is a biological molecule involved in replication of cells. CDC7 is implicated in several cancer types for which there is an unmet need for additional treatments. Drugs which inhibit CDC7 activity (CDC7 inhibitors) have been widely trialled by the pharmaceutical industry. However, despite many attempts over 15 years, CDC7 inhibitor drug candidates have not been successful in clinical trials. We believe that a deeper understanding of the effects of CDC7 inhibitors on cancer biology and human genetics will greatly increase the chances of successful CDC7 inhibitor development, as it will enable us to more confidently select which patients can benefit from CDC7 inhibitors. Turbine’s Simulated Cell technology is a computational approach that is being used to understand which genetic differences make a tumour sensitive to a novel CDC7 inhibitor drug candidate that is being developed by CRH, the commercialisation and development arm of Cancer Research UK. The project now plans to compare our findings with the patient data available in the S:CORT database to help inform future experiments and patient selection strategies for CRH’s novel CDC7 inhibitors. Once we better understand this relationship between CRH’s novel CDC7 inhibitors and the genetics of tumours, we hope to use that knowledge to test the therapeutic effect of this new compound in patients that are most likely to benefit from it.
As systemic treatment options for colorectal cancer have grown, medical oncologists are increasingly confronting decisions about which regimen is optimal for a patient with advanced colorectal cancer. Unfortunately, there are limited biomarkers, patterns in a patients data or tissue, available at present to help clinicians guide patients through treatment decisions where uncertainty in the medical literature exists. Valar Labs is a Stanford-based start-up company that seeks to use artificial intelligence to identify features which are not readily identifiable by the human eye in routine, diagnostic microscope slides (pathology) of tumors, but are nevertheless associated with cancer treatment responses. This project aims to use the pathology images that are collected under S:CORT to attempt to identify biomarkers for response to standard of care treatments from the digital pathology images that form part of the S:CORT cohort using Valar’s AI-enabled digital pathology platform.
The differing subtypes of Bowel (colorectal) cancer, defined by molecular signatures, exhibit different clinical behaviour including response to treatment and prognosis. Traditionally, molecular profiling techniques have been used to diagnose a patient’s particular sub-type of colorectal cancer but this is complex and time-consuming. However, Ground Truth Labs have found that artificial intelligence (AI) can accurately predict these signatures, specifically Consensus Molecular Subtypes (CMS), using digital pathology images, offering a faster, more efficient alternative.
The goal of this project is to create a clinical-grade AI algorithm to predict these molecular subtypes and assess their correlation with patient outcomes, starting with a CMS predictor.
The expansion of computer science and artificial intelligence approaches, such as machine learning, hold great promise for the potential to discover causes of cancer and develop new treatments. Machine learning algorithms can use clinical data to learn and make predictions about the biology of cancer. insitro is using machine learning to improve the understanding of colorectal cancer, subtypes of the disease and possible new treatments. This includes using these techniques to study data arising from clinical trials and NHS care.
In this project Insitro will use the S:CORT Data to find new ways of selecting patients likely to respond to existing therapies, find new targets for treatments, or new uses of existing treatments. To achieve these goals, Insitro will:
The outcomes from this project have the potential to benefit patients by increasing the precision of treatment decisions so that more patients receive the treatment that is most likely to work for them.